All content on this site is intended for healthcare professionals only. By acknowledging this message and accessing the information on this website you are confirming that you are a healthcare professional. If you are a patient or carer, please visit Know AML.
The AML Hub website uses a third-party service provided by Google that dynamically translates web content. Translations are machine generated, so may not be an exact or complete translation, and the AML Hub cannot guarantee the accuracy of translated content. The AML Hub and its employees will not be liable for any direct, indirect, or consequential damages (even if foreseeable) resulting from use of the Google Translate feature. For further support with Google Translate, visit Google Translate Help.
The AML Hub is an independent medical education platform, sponsored by Daiichi Sankyo, Johnson & Johnson, Syndax, Thermo Fisher Scientific, Kura Oncology, and AbbVie. Funders are allowed no direct influence on our content. The levels of sponsorship listed are reflective of the amount of funding given. View funders.
Now you can support HCPs in making informed decisions for their patients
Your contribution helps us continuously deliver expertly curated content to HCPs worldwide. You will also have the opportunity to make a content suggestion for consideration and receive updates on the impact contributions are making to our content.
Find out more
Create an account to access:
Bookmark & personalize site content
Receive alerts for new content in your areas of interest
View AML content recommended for you
Next-generation sequencing-based analysis in patients from the phase III ADMIRAL trial
The recent phase III ADMIRAL trial (NCT02421939) of gilteritinib, an oral, highly specific type 1 FMS-like tyrosine kinase inhibitor, demonstrated clinical superiority to salvage chemotherapy,1 leading to its approval in the U.S. and Europe for relapsed or refractory (R/R) FLT3-mutated acute myeloid leukemia (AML). However, treatment resistance has limited the duration of response to gilteritinib, thought to be due to both on- and off-target mechanisms.1 Catherine Smith and colleagues1 reported results in Blood Advances, on a next-generation sequencing (NGS)-based analysis from the patient cohort in the above-mentioned trial, which aimed to:
- assess the mutational spectrum before and after gilteritinib treatment
- analyze the relationship between mutational spectrum and treatment outcomes
- determine the impact of FLT3-ITD allelic ratio and length on treatment response and overall survival (OS)
We summarize the findings below.
Study design
Blood or bone marrow samples from the patient cohort in the ADMIRAL trial were collected during study entry and at relapse. Baseline co-mutations in 37 recurrently mutated genes were analyzed in 361 evaluable patients, along with full exon sequencing for FLT3 and mutational hotspots for all other genes. Allelic ratio and gene length were measured in 335 evaluable patients, with median ratio calculated at 0.77. Allelic ratios ≥0.77 were classified as high, and <0.77 as low. Median FLT3-ITD length was 51 base pairs (bp), and patient outcomes were assessed at FLT3-ITD length >51 bp and ≤51 bp.
Results
The patient cohort was categorized into gene subgroups, as shown in Table 1.
Table 1. Gene subgroups within the patient cohort*
|
*Adapted from Smith et al.1 |
|
|
Gene subgroup†, % |
Patients (N = 361) |
|---|---|
|
DNA methylation/hydroxymethylation |
41.2 |
|
Transcription factors/regulators |
26.3 |
|
Chromatin-spliceosome-other |
17.4 |
|
RTK-Ras signalling |
7.8 |
|
TP53-aneuploidy |
3.6 |
|
NPM1 |
47.9 |
|
DNMT3A |
31.9 |
|
DNMT3A/NPM1 |
23.8 |
|
WT1 |
18.0 |
|
IDH1/IDH2 |
15.5 |
Ras/MAPK pathway gene mutations, which are reportedly associated with gilteritinib resistance, occurred in 6.9% of patients (n = 25/361). No associations between co-mutations and baseline characteristics (age, sex, Eastern Cooperative Oncology Group performance status, cytogenetic risk, or prior treatment with a FLT3 inhibitor) were found.
Impact of baseline co-mutations on response outcomes
Across all gene subgroups except TP53-aneuploidy, rates of complete remission (CR) with full or partial hematologic recovery (CR/CRh) before hematopoietic stem cell transplantation (HSCT) were higher in the gilteritinib arm than in the salvage chemotherapy (SC) arm; however, the TP53-aneuploidy category was small (n = 13). Responses to gilteritinib and SC in each of the gene subgroups are shown in Figure 1.
Figure 1. Response rates to the gilteritinib and SC arms across gene subgroups*
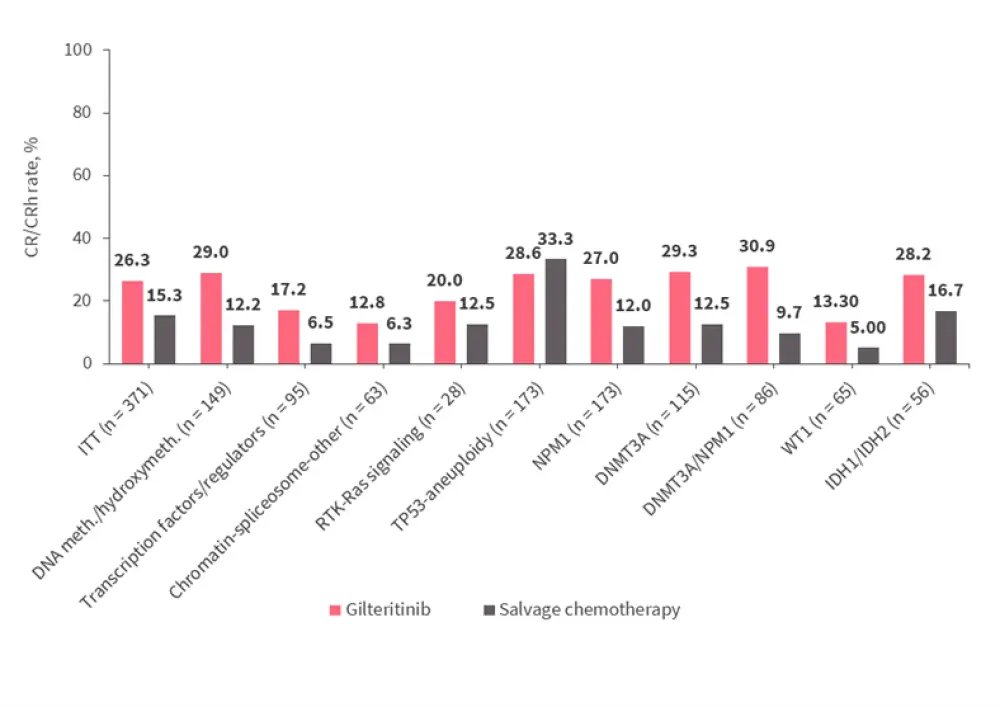
CR, complete remission; CRh, CR with partial hematologic recovery; DNA meth./hydroxymeth., DNA methylation/hydroxymethylation; ITT, intention-to-treat population; SC, salvage chemotherapy.
*Adapted from Smith et al.1
Impact of baseline co-mutations on survival outcomes
Patients treated with gilteritinib had improved median OS compared to those treated with SC, among the following gene categories:
- NPM1-mutated
- DNA methylation/hydroxymethylation
- Transcription factor
- Co-mutated DNMT3A + WT1
- Dual-mutated DNMT3A + NPM1 (most pronounced survival advantage)
Survival with the gilteritinib treatment was similar in co-mutated and wildtype NPM1, DNMT3A, and WT1 subgroups. Survival in the SC arm was reduced in co-mutated NPM1, and WT1 subgroups compared to wildtype NPM1 or WT-1 subgroups, but was similar in co-mutated and wildtype DNMT3A subgroups.
Impact of FLT3-ITD gene length and number of FLT3-ITD mutations on clinical outcomes
Table 2 shows the survival rates for patients with single and multiple FLT3-ITD mutations. A total of 280 patients had a single FLT3-ITD mutation, whereas 55 patients had multiple FLT3-ITD mutations with co-mutation profiles similar across treatment arms.
Table 2. Median OS rates according to FLT3-ITD mutation length and multiple mutations*
|
bp, base pairs; CR, complete remission; CRh, CR with partial hematologic recovery; ITD, internal tandem duplication; OS, overall survival; SC, salvage chemotherapy. |
|||
|
Outcome |
Gilteritinib arm |
SC arm |
HR |
|---|---|---|---|
|
Patients with single FTL3-ITD mutation at baseline, n |
189 |
91 |
— |
|
Median OS based on FTL3-ITD mutation length, months |
|||
|
≤51 bp |
8.9 |
6.1 |
0.807 |
|
>51 bp |
10.4 |
6.0 |
0.480 |
|
Patients with multiple FTL3-ITD mutations at baseline, n |
33 |
22 |
— |
|
Median OS, months |
8.3 |
3.5 |
0.624 |
|
Rates of pretransplant CR/CRh based on FLT3-ITD allelic ratio, % |
|||
|
Low FLT3-ITD ratio (<0.77) |
32.7 |
18.9 |
— |
|
High FLT3-ITD ratio (≥0.77) |
21.1 |
10.0 |
— |
|
Median OS based on FLT3-ITD allelic ratio, months |
|||
|
Low FLT3-ITD ratio (<0.77) |
10.6 |
6.9 |
0.795 |
|
High FLT3-ITD ratio (≥0.77) |
7.1 |
4.3 |
0.49 |
- Improved survival was seen in the gilteritinib treatment arm compared with the SC treatment arm, both in patients with FTL3-ITD mutation length ≤51bp and >51bp
- Mutation length did not appear to impact the survival benefit observed with gilteritinib
- The survival benefit with gilteritinib was also seen in patients with multiple FLT3 mutations, and, moreover, the presence of multiple FLT3 mutations did not affect the response to gilteritinib
- Patients who received gilteritinib and had a gene length >51 bp had the longest median OS
- Presence of multiple mutations did not appear to have a negative impact on survival when compared to short or long ITD subgroups
- Within the SC arm, survival was similar regardless of gene length, although reduced when multiple mutations where present
Impact of allelic ratio on clinical outcomes
Assessment of FLT3-ITD allelic ratio was performed in 335 patients, who were classified as having either a high (≥0.77; n = 169) or low (<0.77; n= 166) ratio. The CR/CRh and survival rates for both treatment groups are shown in Table 2.
- Gilteritinib treatment was associated with improved survival across patients with high and low allelic ratios
- In both treatment groups, survival was reduced in patients with a high allelic ratio
Mutation profiles of patients who relapsed on gilteritinib
Of the 247 patients on gilteritinib therapy, 75 who achieved composite CR (CRc) subsequently relapsed, and median time from first dose to relapse was 4.6 months. Mutation profiles of the 40 patients with available samples for analysis at baseline and relapse are shown in Figure 2. Overall, 27 of these patients had new gene mutations at relapse, with 18 acquiring new mutations in the Ras/MAPK gene pathway.
Figure 2. Mutation profiles of patients who relapsed on gilteritinib treatment*
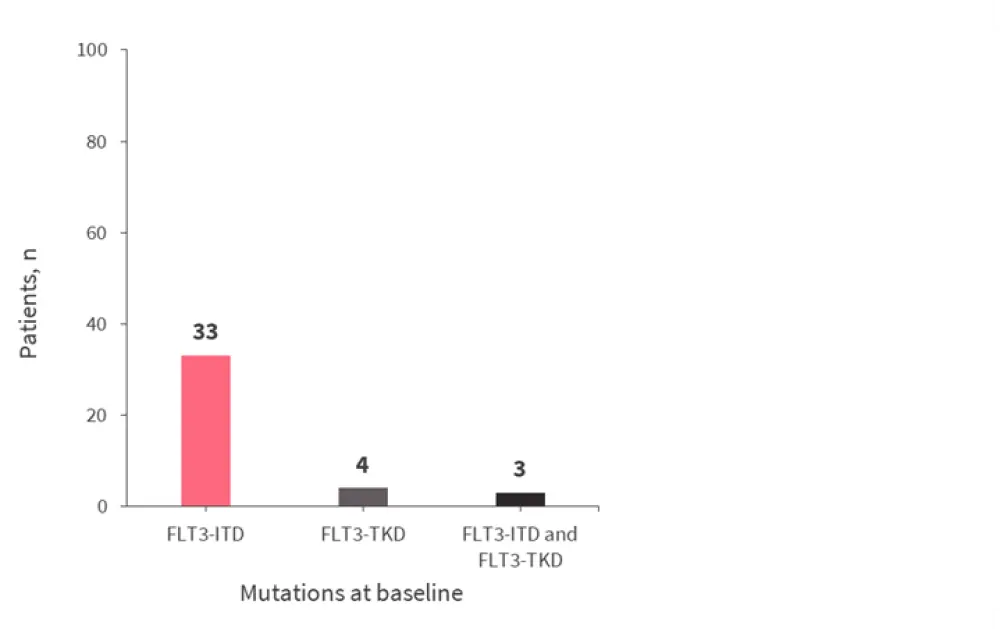
*Adapted from Smith et al.1
Response outcomes in patients with mutated Ras/MAPK pathway genes at baseline
Ras/MAPK gene mutations were present in 25 of 361 patients with available baseline samples (gilteritinib, n = 18; SC, n = 7).
- CRc rate prior to HSCT in patients treated with gilteritinib with baseline Ras/MAPK mutations was 33.3%; median duration of CRc was not estimable
- CR/CRh rate in patients with baseline Ras/MAPK mutations was 22.2% for those treated with gilteritinib and 14.3% for those treated with SC
Conclusion
Overall, patients with a low FLT3 allelic ratio and gene length >51 bp who received gilteritinib had improved remission and survival rates compared to those who received SC. The study1 also confirmed the highly selective nature of gilteritinib and the significant role of the FLT3 pathway, while highlighting the potential of FLT3-targeted therapies in altering mutation profiles over time.
References
Please indicate your level of agreement with the following statements:
The content was clear and easy to understand
The content addressed the learning objectives
The content was relevant to my practice
I will change my clinical practice as a result of this content
Your opinion matters
Which AML-related topic do you currently need the most practical guidance on?

